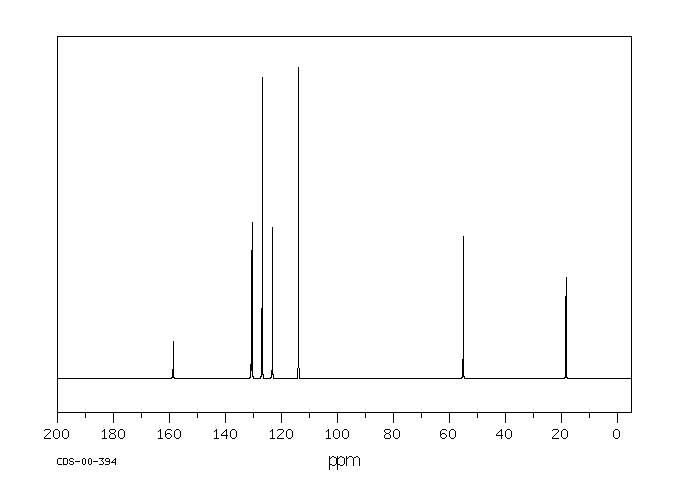
Whether the Der p 2 fold is a new fold or a distant relative of the α-amylase inhibitor or the immunoglobulin superfamily remains to be determined. However, a crucial element of the immunoglobulin superfamily is the Greek-key motif, and Der p 2 is lacking the strand that would define a Greek-key. Strand 4 of the immunoglobulin superfamily has been replaced with an extended loop that comprises residues 59–78 of Der p 2. An examination of other strand-loop-strand motifs in the data base revealed that the two-dimensional model of Der p 2 appears similar to that of the immunoglobulin superfamily, also shown in Fig. The arrangement of secondary structural elements in α-amylase inhibitor is clearly unlike that of Der p 2. The alignment of the sheets relative to the preliminary model of Der p 2 is shown in Fig. Only a single potential structural homologue, the α-amylase inhibitor (1hoe.pdb), was found. Since there appear to be two three-stranded anti-parallel ॆ-pleated sheets, a search of several structural data bases was initiated. A value of 1 indicates that the amide proton of that residue remained protonated for greater than 1 week after exchange into D 2O buffer. Residues that could not be found in the HNHA or were too degenerate to measure accurately are indicated with a zero bar height. These ranges are indicated by dashed lines.  .pdf+-+SumatraPDF_2012-12-31_06-06-40.png)
ॆ-sheet residues typically have coupling constants of 8–10 Hz, and α-helices typically have coupling constants of 4–7 Hz. The bar height indicates to J HN-HA coupling constant for that residue. Blanks indicate the absence of a cross-peak or a residue that was too degenerate to determine unambiguously. The bar height indicates the intensity, on an arbitrary scale, of the H α(i) − H N(i+1) NOESY cross-peak in the 15N-NOESY-HSQC, CHH-NOESY, and CN-NOESY. The bar height indicates the intensity, on an arbitrary scale, of the H N(i) − H N(i+1) NOESY cross-peak in the 15N-NOESY-HSQC. The bars indicate the predicted secondary structure for each residue based on the chemical shift of the C α, C ॆ, CO, and H α. 
Isolation of r Der p 2 (D1S)įigure 3 Secondary structure of rDer p 2 (D1S). Therefore, the disulfide bonds present in the final folded recombinant molecule most probably were formed during synthesis and not during the isolation procedure. Note that at no time did the extraction or refolding buffers contain any disulfide exchange reagents, e.g.mercaptoethanol, dithiothreitol, and glutathione. After dialysis, NaCl was added to 100 m mconcentration, and the sample was centrifuged (GS-34 rotor, 15,000 rpm for 20 min). To be more accurate, let’s mention that it is the ratio of the protons behind each signal. The pellet was dissolved in one-fifth of the original culture volume of 6 m guanidine-HCl and dialyzed against 20 m mTris, pH 8.5, 1 m m EDTA without stirring overnight at room temperature, followed by a buffer change and an additional 4 h of dialysis. The integration in NMR tells us the number of protons represented by a given signal. The sonicate was centrifuged (GS-34 rotor, 12,000 rpm for 20 min), and the supernatant was discarded. The cell pellet was frozen for 3–4 h at −20 ☌ and subsequently thawed and resuspended to one-twentieth of the original culture volume in TE (100 m m Tris, pH 8.5, 10 m m EDTA) and sonicated for 2 min on ice. Glycobiology and Extracellular Matrices.Starcuk, Z., Pcek, L., Fiala, R., and Starcuk Jr., Z., 1988, J. Plateau, P., Dumas, C., and Gueron, M., 1983, J. A., Loaiza, F., Valentine, K., and Warren, W. R., 1989, Fourier Transforms in Optical, NMR, and Mass Spectrometry: A User’s Handbook,” Elsevier, Amsterdam. Hochmann, J., and Kellerhals, H., 1980, J. Gutow, J., McCoy, M., Sparo, F., and Warren, W. This process is experimental and the keywords may be updated as the learning algorithm improves.Īrata, Y., and Ozawa, H., 1976, J. These keywords were added by machine and not by the authors. Even in D 2O, the residual HDO signal can be large enough to obscure signals at or near the HDO peak. Although much can be learned about the nonexchangeable protons from FT/NMR conducted in D 2O solution, exchangeable protons (e.g., base-pair protons in RNA and DNA) can obviously be observed only in H 2O. Fourier transform NMR in dilute aqueous solutions is essential for the analysis of the solution structure and dynamics of biological molecules (e.g., proteins, carbohydrates, phospholipids, RNA, DNA), drugs, and a wide variety of polar molecules (Wüthrich, 1976 Wüthrich,1986 Campbell-Burke and Shulman, 1987 Brindle and Campbell, 1987 Shulman et al., 1979 Radda and Seeley, 1979 Kuchel et al., 1984 Rabenstein, 1984 Scott and Baxter, 1981 Roberts and Jandetzky, 1981).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |



 RSS Feed
RSS Feed
